教程3:Cell Classification(细胞分类)¶
目标:展示如何基于 GridScore 与放电率统计进行细胞分类,并输出分类结果与可视化。
结构:
任务概览
计算 GridScore
分类方法示例
输出与可视化
Note
本教程以 examples/cell_classification 中脚本为参考,偏重示例与流程。
若需要完整 ASA pipeline,请先阅读 教程1:ASA Pipeline:实验数据解码全流程。
0. 任务概览¶
细胞分类通常基于以下信息:
GridScore:衡量放电场是否呈网格结构
放电率与空间选择性:用于筛选候选细胞
模块划分:通过自相关特征聚类识别不同网格模块
本教程展示两条实践路径:
单细胞 GridScore 评分与可视化
基于自相关特征的模块识别(分类)
1. 计算 GridScore¶
示例脚本:examples/cell_classification/grid_score.py
关键步骤:
embed_spike_trains将 spike 事件嵌入为(T, N)矩阵compute_rate_map_from_binned生成 rate mapcompute_2d_autocorrelation计算自相关GridnessAnalyzer.compute_gridness_score得到 GridScore
[ ]:
from canns.analyzer import data
from canns.analyzer.data.cell_classification import (
GridnessAnalyzer,
compute_2d_autocorrelation,
compute_rate_map_from_binned,
plot_autocorrelogram,
plot_rate_map,
)
from canns.data.loaders import load_grid_data, load_left_right_npz
grid_data = load_grid_data()
cfg = data.SpikeEmbeddingConfig(smooth=False, speed_filter=True, min_speed=0.0)
spikes, xx, yy, _tt = data.embed_spike_trains(grid_data, config=cfg)
analyzer = GridnessAnalyzer()
neuron_id = 69
rate_map, *_ = compute_rate_map_from_binned(xx, yy, spikes[:, neuron_id], bins=35)
autocorr = compute_2d_autocorrelation(rate_map)
result = analyzer.compute_gridness_score(autocorr)
print(f"neuron {neuron_id}\nscore: {result.score}\nspacing: {result.spacing}\norientation: {result.orientation}")
# Visualize rate map + autocorrelogram
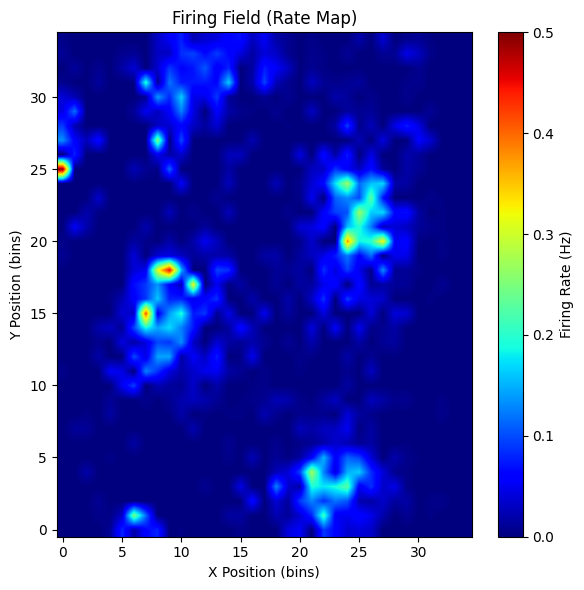
plot_rate_map(rate_map, show=True)
plot_autocorrelogram(autocorr, gridness_score=result.score, show=True)
Loaded grid_1: 172 neurons
Position data available: 238700 time points
neuron 69
score: 0.8761650577011183
spacing: [18.43908891 17.08800749 18.11077028]
orientation: [-40.60129465 20.55604522 83.65980825]
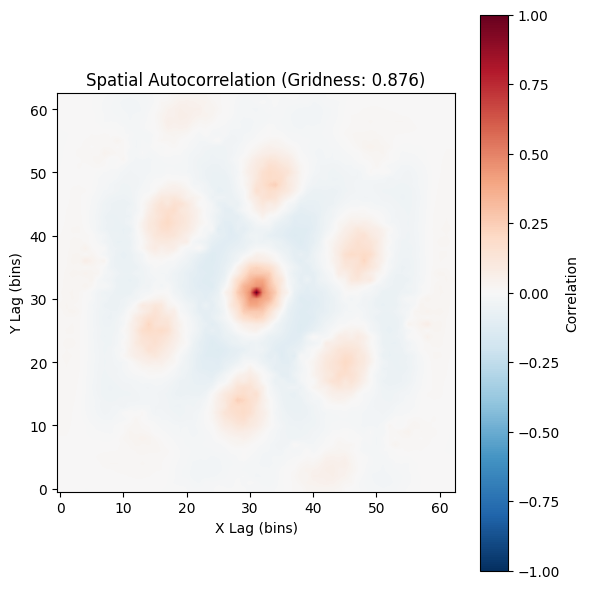
(<Figure size 600x600 with 2 Axes>,
<Axes: title={'center': 'Spatial Autocorrelation (Gridness: 0.876)'}, xlabel='X Lag (bins)', ylabel='Y Lag (bins)'>)
2. 分类方法示例¶
示例脚本:examples/cell_classification/try_classification_methods.py
该示例使用自相关图作为特征,采用 Leiden 聚类识别网格模块。 核心函数:
identify_grid_modules_and_stats:输出模块划分与统计信息
[8]:
from canns.analyzer.data.cell_classification import (
GridnessAnalyzer,
identify_grid_modules_and_stats,
compute_rate_map_from_binned,
compute_2d_autocorrelation,
)
import numpy as np
grid_data = load_left_right_npz(
session_id="28304_1",
filename="28304_1_ASA_mec_full_cm.npz",
)
n_units = spikes.shape[1]
autocorrs = []
for nid in range(n_units):
rate_map, *_ = compute_rate_map_from_binned(xx, yy, spikes[:, nid], bins=35)
autocorr = compute_2d_autocorrelation(rate_map)
autocorrs.append(autocorr.astype(np.float32, copy=False))
autocorrs = np.stack(autocorrs, axis=0)
analyzer = GridnessAnalyzer()
out = identify_grid_modules_and_stats(
autocorrs,
gridness_analyzer=analyzer,
k=30,
resolution=1.0,
score_thr=0.3,
consistency_thr=0.5,
min_cells=10,
merge_corr_thr=0.7,
metric='manhattan',
)
print(out['n_grid_cells'], out['n_modules'])
106 1
/Users/sichaohe/Documents/GitHub/canns/src/canns/analyzer/data/cell_classification/utils/image_processing.py:284: FutureWarning: `binary_dilation` is deprecated since version 0.26 and will be removed in version 0.28. Use `skimage.morphology.dilation` instead. Note the lack of mirroring for non-symmetric footprints (see docstring notes).
dilated = morphology.binary_dilation(image, footprint=footprint)
3. 输出与可视化¶
grid_score.py会绘制 rate map、自相关图与 GridScore 直方图try_classification_methods.py会保存grid_modules_demo_output.npz
建议:
先用小批量神经元验证参数,再扩大到全体
适当调整
bins/score_thr/consistency_thr以匹配数据质量若需与 ASA pipeline 联动,可将
spikes与x/y/t与解码结果结合分析