如何分析实验数据?¶
目标: 了解 ASA pipeline 的核心步骤,并能用 Python 跑通 TDA → Decode → CohoMap/CohoSpace/PathCompare 等分析。
预计阅读时间: 12 分钟
.. note:: 如需深入了解拓扑解码与神经群体流形,可参阅 :cite:p:`Vaupel2023Duality,Gardner2022Toroidal`。ASA pipeline 概览¶
ASA 将实验记录统一为 ASA .npz (包含 spike, x, y, t ),核心流程如下:
embed_spike_trains—— 将 spike times 嵌入为(T, N)活动矩阵tda_vis—— 持续同调与 barcodedecode_circular_coordinates_multi—— 解码相位轨迹(coords/coordsbox/times_box)plot_cohomap_multi—— 相位投影到真实轨迹plot_cohospace_*_1d/plot_cohospace_*_2d/compute_cohoscore_1d/compute_cohoscore_2d—— 神经元相位选择性plot_path_compare_1d/plot_path_compare_2d—— 真实轨迹 vs 解码轨迹compute_fr_heatmap_matrix/compute_frm—— FR/FRM
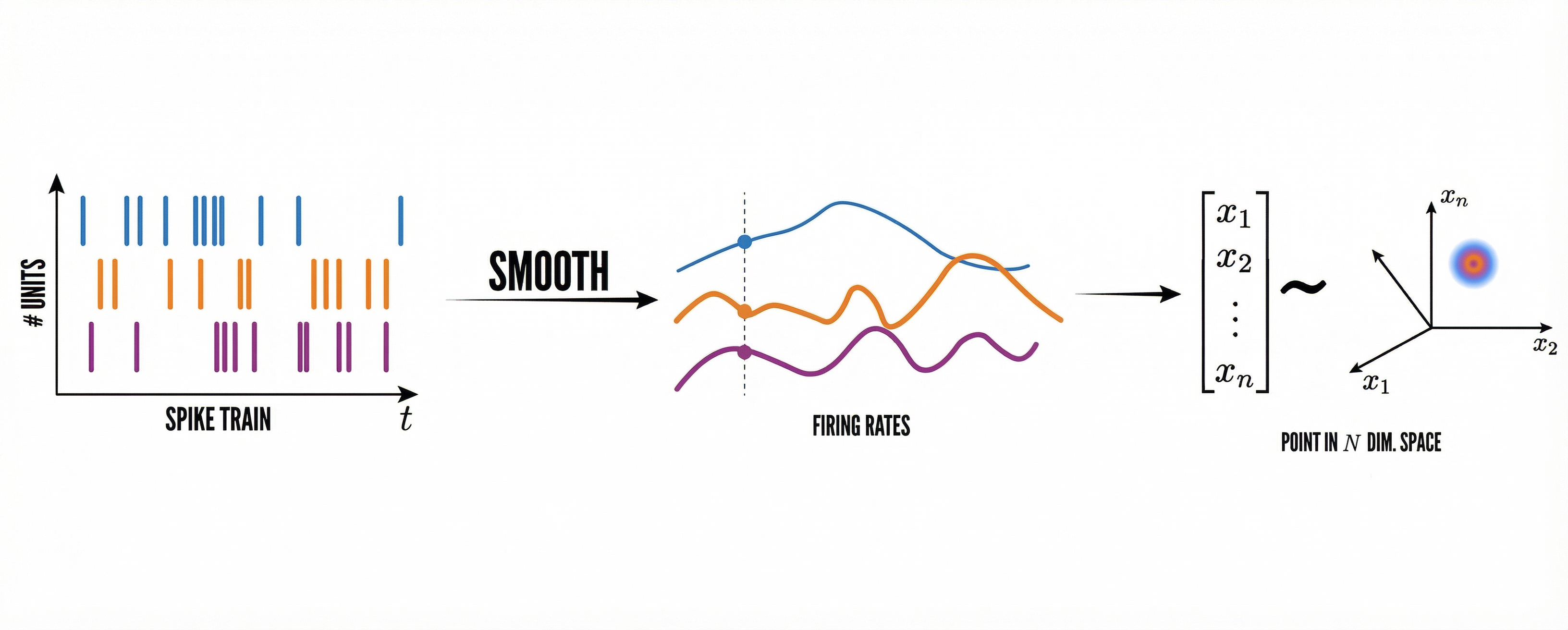
Spike trains 输入¶
示例:内置 grid 数据¶
以下示例使用 ``load_grid_data``(内置网格细胞数据),演示从预处理到 CohoMap 的最小流程。
[1]:
from canns.data.loaders import load_grid_data
grid_data = load_grid_data()
print(f"Keys: {list(grid_data.keys())}")
print(f"t: {grid_data['t'].shape}, x: {grid_data['x'].shape}, y: {grid_data['y'].shape}")
print(f"spike type: {type(grid_data['spike'])}")
Loaded grid_1: 172 neurons
Position data available: 238700 time points
Keys: ['spike', 't', 'x', 'y']
t: (238700,), x: (238700,), y: (238700,)
spike type: <class 'numpy.ndarray'>
步骤 1:嵌入 spike trains(可选)¶
如果 spike 是 list-of-arrays / dict(每个神经元一个 spike times 数组),使用
embed_spike_trains 将其转为 (T, N) 连续矩阵;若已是 (T, N),可跳过。
[2]:
from canns.analyzer import data
spike_cfg = data.SpikeEmbeddingConfig(
smooth=True,
speed_filter=False,
min_speed=2.5,
)
spikes, xx, yy, tt = data.embed_spike_trains(grid_data, config=spike_cfg)
print(f"Embedded spikes: {spikes.shape}")
Embedded spikes: (238700, 172)
步骤 2:TDA + 解码 + CohoMap¶
tda_vis 输出持续同调结果;随后使用 decode_circular_coordinates_multi 获取相位轨迹,
并通过 plot_cohomap_multi 将相位映射到真实轨迹上。
[8]:
from canns.analyzer.visualization import PlotConfigs
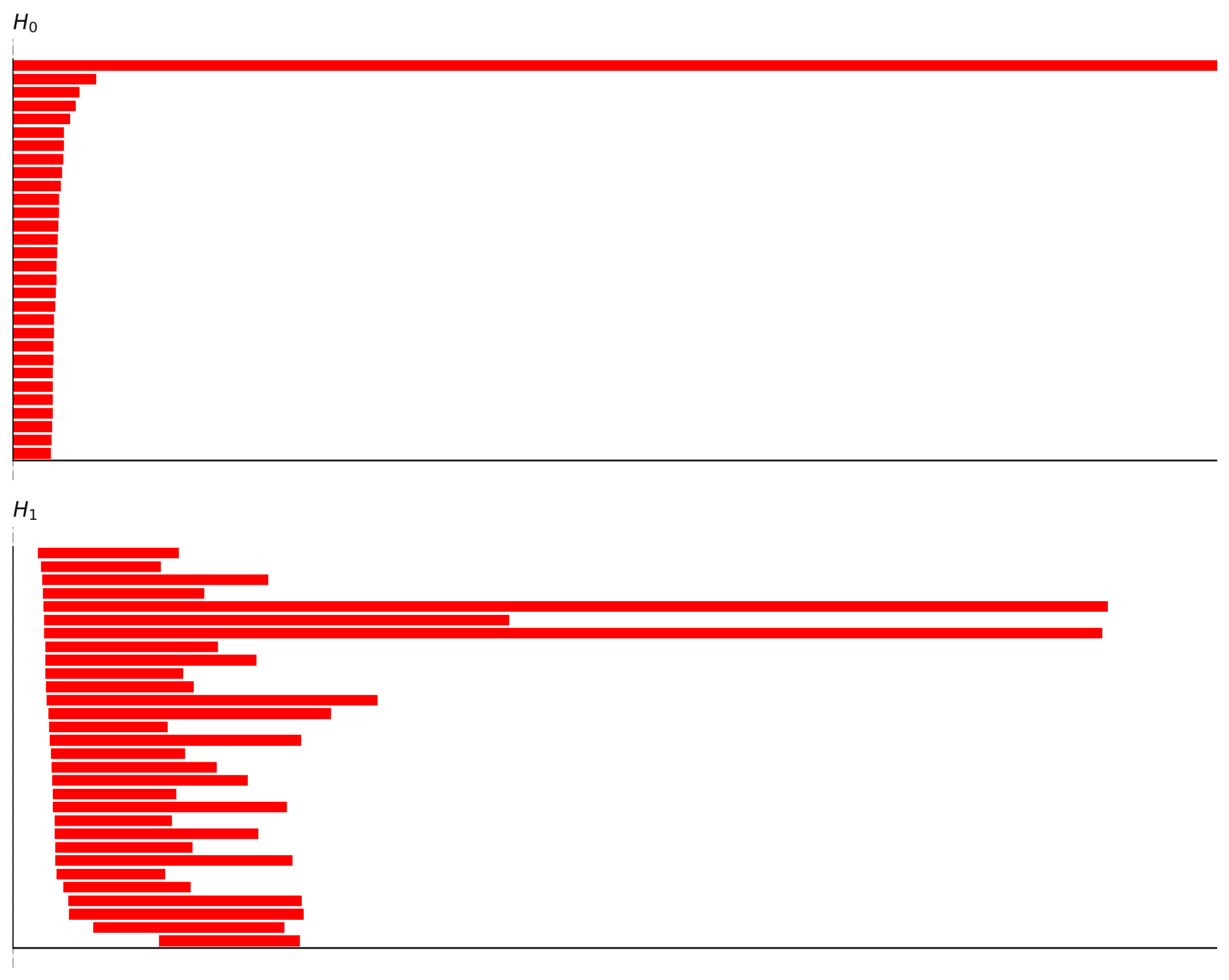
tda_cfg = data.TDAConfig(maxdim=1, do_shuffle=False, show=True, progress_bar=True)
persistence = data.tda_vis(embed_data=spikes, config=tda_cfg)
grid_data_embedded = dict(grid_data)
grid_data_embedded["spike"] = spikes
decoding = data.decode_circular_coordinates_multi(
persistence_result=persistence,
spike_data=grid_data_embedded,
num_circ=2,
)
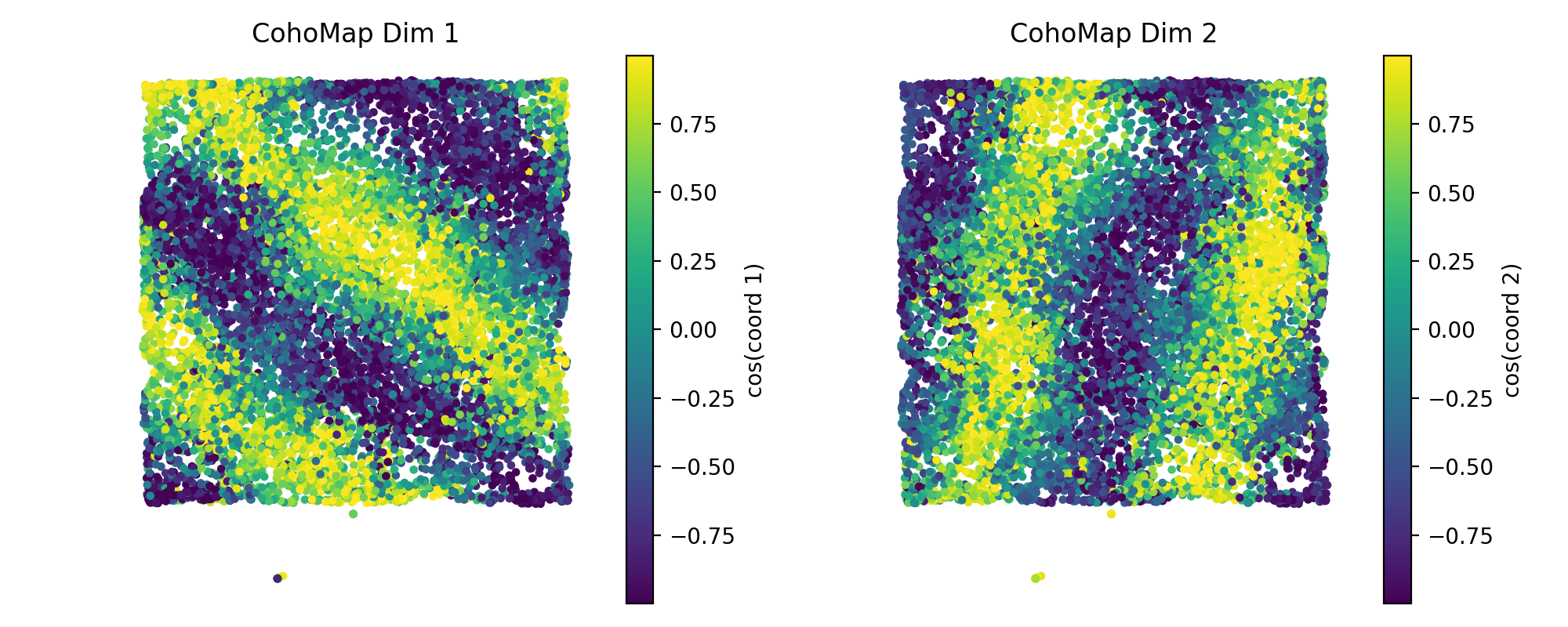
config = PlotConfigs.cohomap(show=True)
fig = data.plot_cohomap_multi(
decoding_result=decoding,
position_data={"x": grid_data["x"], "y": grid_data["y"]},
config=config,
)
Computing persistent homology for real data...
Completed H1 computation: 100%|██████████| 718201/718201 [00:05<00:00, 122566.90columns/s]
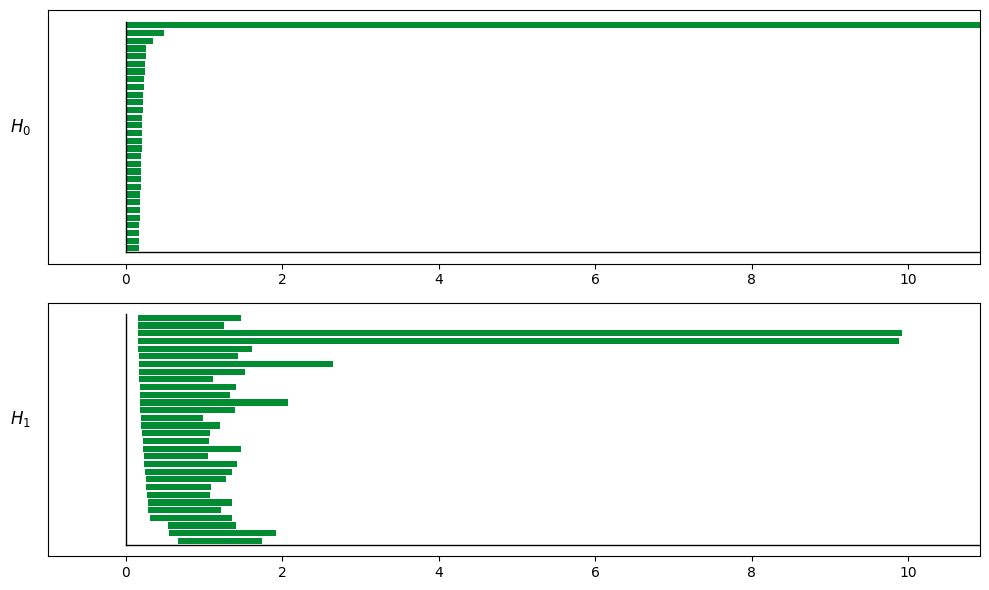
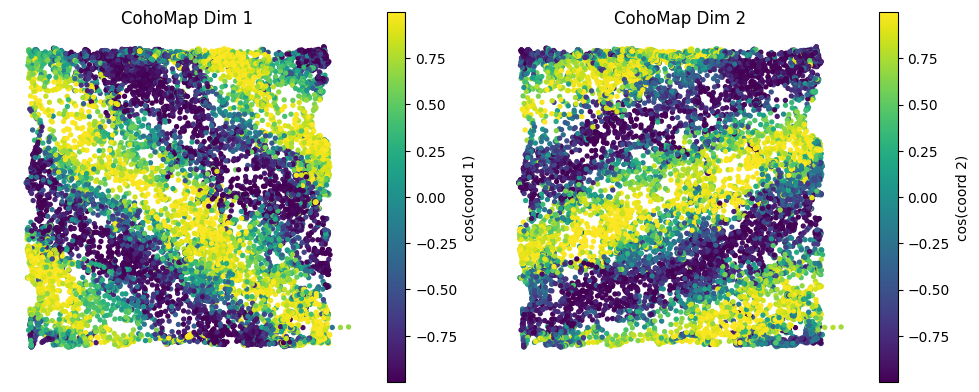
解码结果说明:
coords: 解码相位(T' × K)coordsbox: 用于对齐/绘图的相位(T' × K)times_box:coordsbox对应的时间索引(T')
步骤 3:Path Compare(真实轨迹 vs 解码轨迹)¶
使用 align_coords_to_position_1d / align_coords_to_position_2d 对齐时间索引,再用 plot_path_compare_1d / plot_path_compare_2d 绘制对比。
[6]:
import numpy as np
from canns.analyzer import data
from canns.analyzer.visualization import PlotConfigs
coords = np.asarray(decoding.get("coords"))
_t = np.asarray(grid_data["t"]).ravel()[200:260]
_x = np.asarray(grid_data["x"]).ravel()[200:260]
_y = np.asarray(grid_data["y"]).ravel()[200:260]
t_use, x_use, y_use, coords_use, _ = data.align_coords_to_position_2d(
t_full=_t,
x_full=_x,
y_full=_y,
coords2=coords,
use_box=True,
times_box=decoding.get("times_box", None),
interp_to_full=True,
)
coords_use = data.apply_angle_scale(coords_use, "rad")
plot_config = PlotConfigs.path_compare_2d(show=True)
data.plot_path_compare_2d(x_use, y_use, coords_use, config=plot_config)
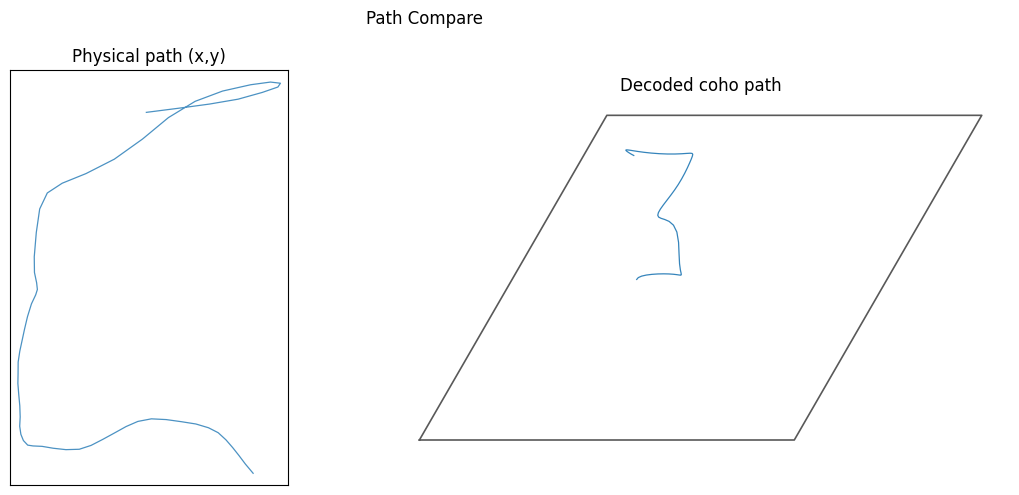
[6]:
(<Figure size 1200x500 with 2 Axes>,
array([<Axes: title={'center': 'Physical path (x,y)'}>,
<Axes: title={'center': 'Decoded coho path'}>], dtype=object))
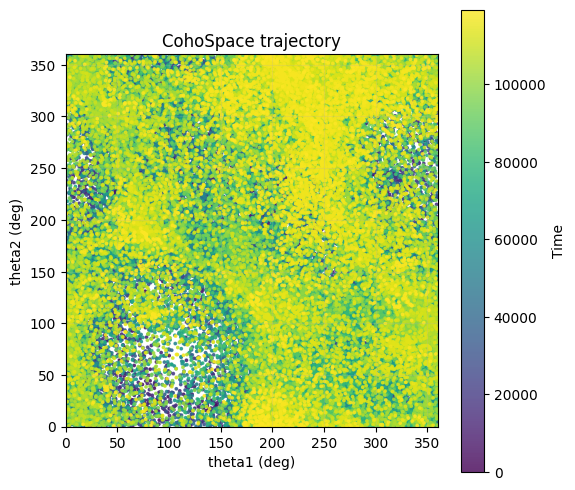
步骤 4:CohoSpace / CohoScore¶
CohoSpace 用于查看神经元在相位空间上的偏好分布。
[7]:
import numpy as np
from canns.analyzer import data
from canns.analyzer.visualization import PlotConfigs
coords = np.asarray(decoding.get("coords"))
coordsbox = np.asarray(decoding.get("coordsbox"))
traj_cfg = PlotConfigs.cohospace_trajectory_2d(show=True)
data.plot_cohospace_trajectory_2d(
coords=coords[:, :2],
times=None,
subsample=2,
config=traj_cfg,
)
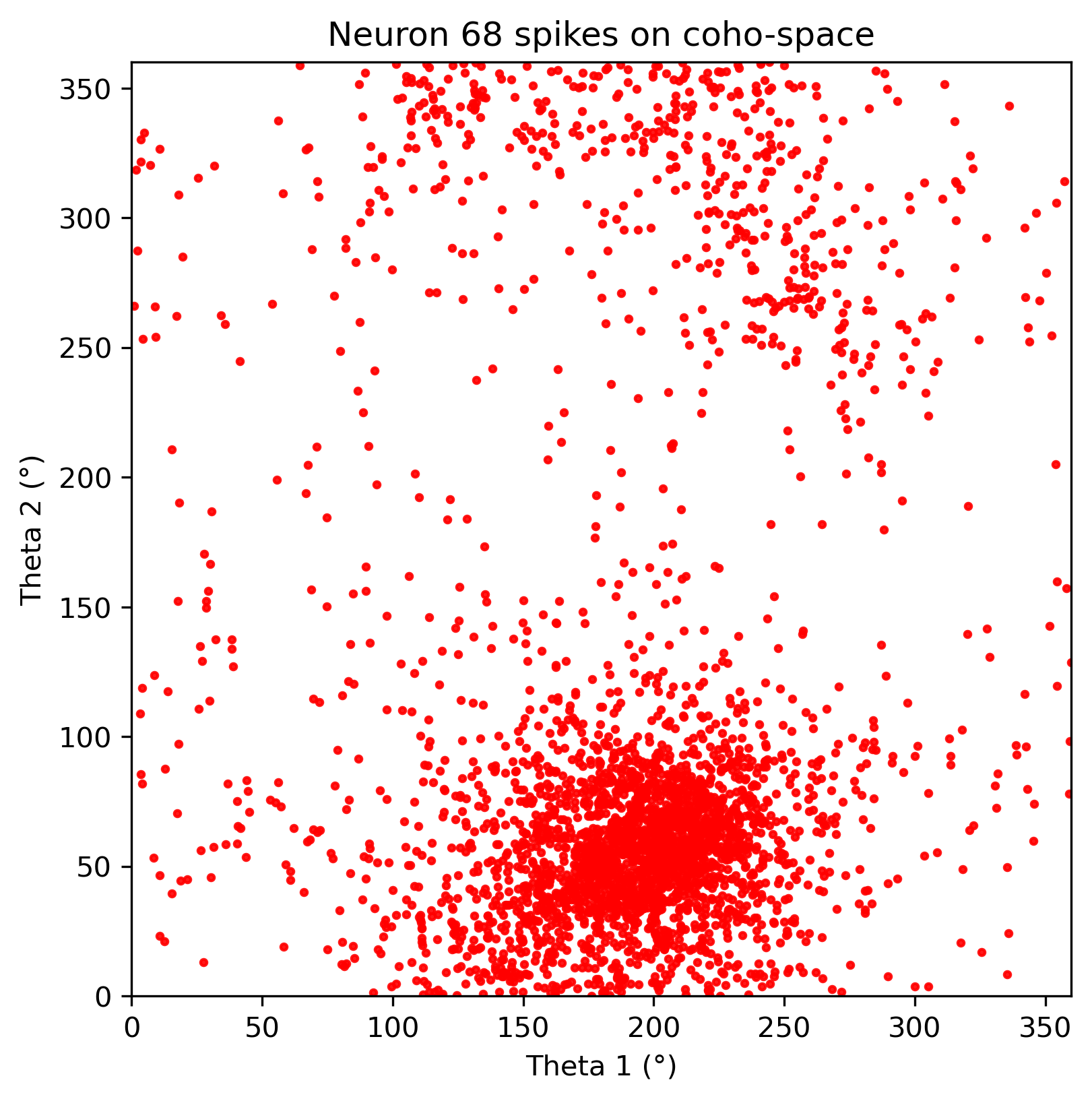
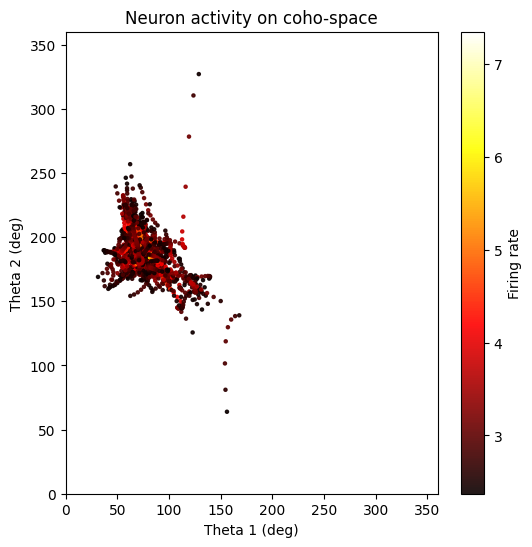
neuron_cfg = PlotConfigs.cohospace_neuron_2d(show=True)
fig = data.plot_cohospace_neuron_2d(
coords=coordsbox[:, :2],
activity=spikes,
neuron_id=130,
mode="fr",
top_percent=1,
config=neuron_cfg,
)
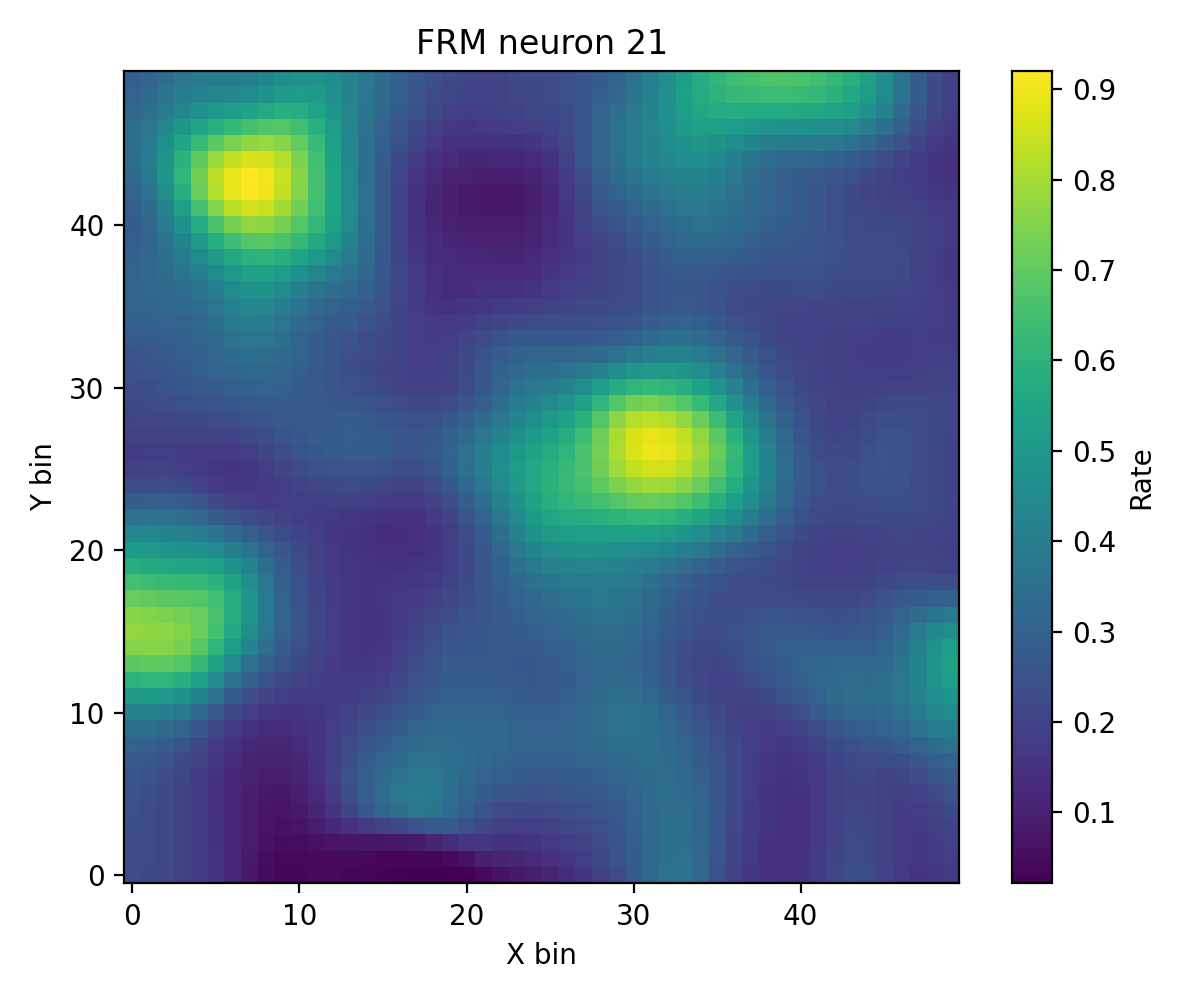
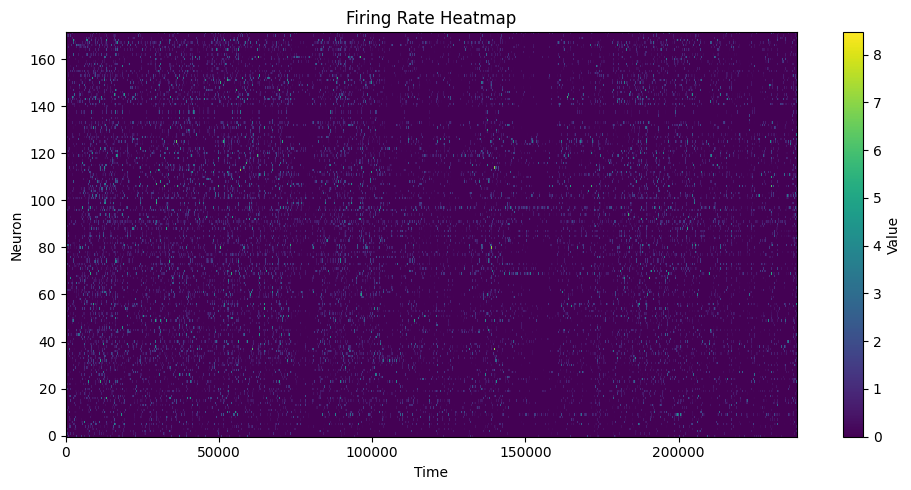
步骤 5:FR Heatmap / FRM¶
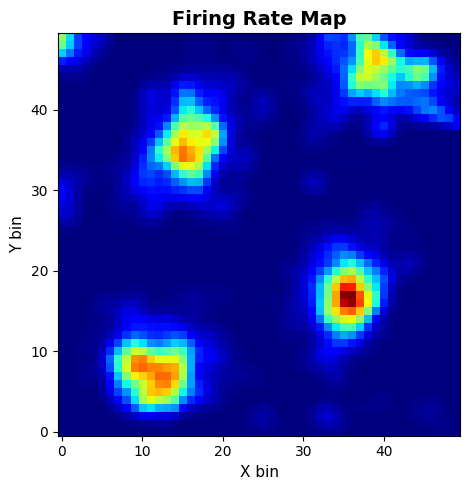
快速检查群体活动(FR Heatmap)以及单神经元空间放电图(FRM)。
[10]:
from pathlib import Path
from canns.analyzer.visualization import PlotConfigs
heatmap = data.compute_fr_heatmap_matrix(spikes, transpose=True)
heatmap_cfg = PlotConfigs.fr_heatmap(show=True)
data.save_fr_heatmap_png(heatmap, config=heatmap_cfg)
frm_res = data.compute_frm(
spikes,
grid_data["x"],
grid_data["y"],
neuron_id=130,
bins=50,
min_occupancy=1,
smoothing=True,
sigma=1.0,
nan_for_empty=True,
)
frm_cfg = PlotConfigs.frm(show=True)
data.plot_frm(frm_res.frm, config=frm_cfg, dpi=200)
数据格式与注意事项¶
ASA 输入格式:
spike,x,y,t``(``t与spike时间单位一致)spike支持 dict / list-of-arrays /(T, N)矩阵如果启用
speed_filter,需要x/y与t对齐
后续步骤¶
继续学习:
核心概念:分析方法 —— 了解 ASA pipeline 与 TDA 的设计思想
完整 API 参考:数据分析器 ——
canns.analyzer.data的所有接口TUI 工作流 —— 交互式 ASA pipeline(后续章节)
Fly ROI 1D bump 拟合示例 ——
examples/experimental_data_analysis/fly_roi_bump_fit.py(roi_bump_fits)