Tutorial 1: ASA GUI End-to-End Analysis Tutorial¶
This tutorial introduces how the ASA GUI (Attractor Structure Analyzer) integrates data preprocessing, TDA, decoding, and visualization from the ASA pipeline into a graphical workflow. You will learn to complete an analysis using the GUI and manage outputs in the results directory.
Note
For an in-depth understanding of analysis principles and parameter selection, please refer to Tutorial 1: ASA Pipeline: End-to-End Decoding of Experimental Data.
Tutorial Objectives¶
Complete an end-to-end analysis using the ASA GUI
Understand input data structure, parameter meanings, and output directory layout
Master common analysis patterns and their dependencies
Target Audience¶
Users who prefer a graphical interface to run the ASA pipeline
Experimental researchers who want to quickly browse analysis results and figures
Experimental teams aiming to reuse TDA/decoding/visualization workflows
Prerequisites¶
CANNs installed (recommended via
pip install -e .orpip install canns)GUI dependencies installed:
pip install canns[gui](includes PySide6)Optional:
pip install qtawesome(for navigation icons)Prepared ASA
.npzdata file
Note
The ASA GUI currently supports only ASA .npz input. Controls related to “Neuron + Trajectory” are reserved but not yet functional.
Launching the ASA GUI¶
Execute in your project environment:
canns-gui
# or
python -m canns.pipeline.asa_gui
Upon launch, the main window appears with a default size of approximately 1200×800, which can be resized according to your screen.
Interface Overview¶
The top of the main window includes:
Preprocess and Analysis navigation buttons
Light / Dark theme toggle
Help button (opens quick usage instructions)
Chinese / English language toggle
The Preprocess page handles input configuration, preprocessing parameters, and logs. The Analysis page manages analysis parameters, result previews, and file output.
Interface Elements Inventory¶
Preprocess Page¶
Input / Mode |
Fixed as |
Preset |
Preset templates: |
ASA file |
Drag and drop or click Browse to select a |
Preprocess / Method |
Choose |
Embedding Parameters |
|
Pre-classification |
Reserved options ( |
Run / Stop / Progress |
Start or terminate preprocessing and display a progress bar. |
Logs |
Display preprocessing logs. |
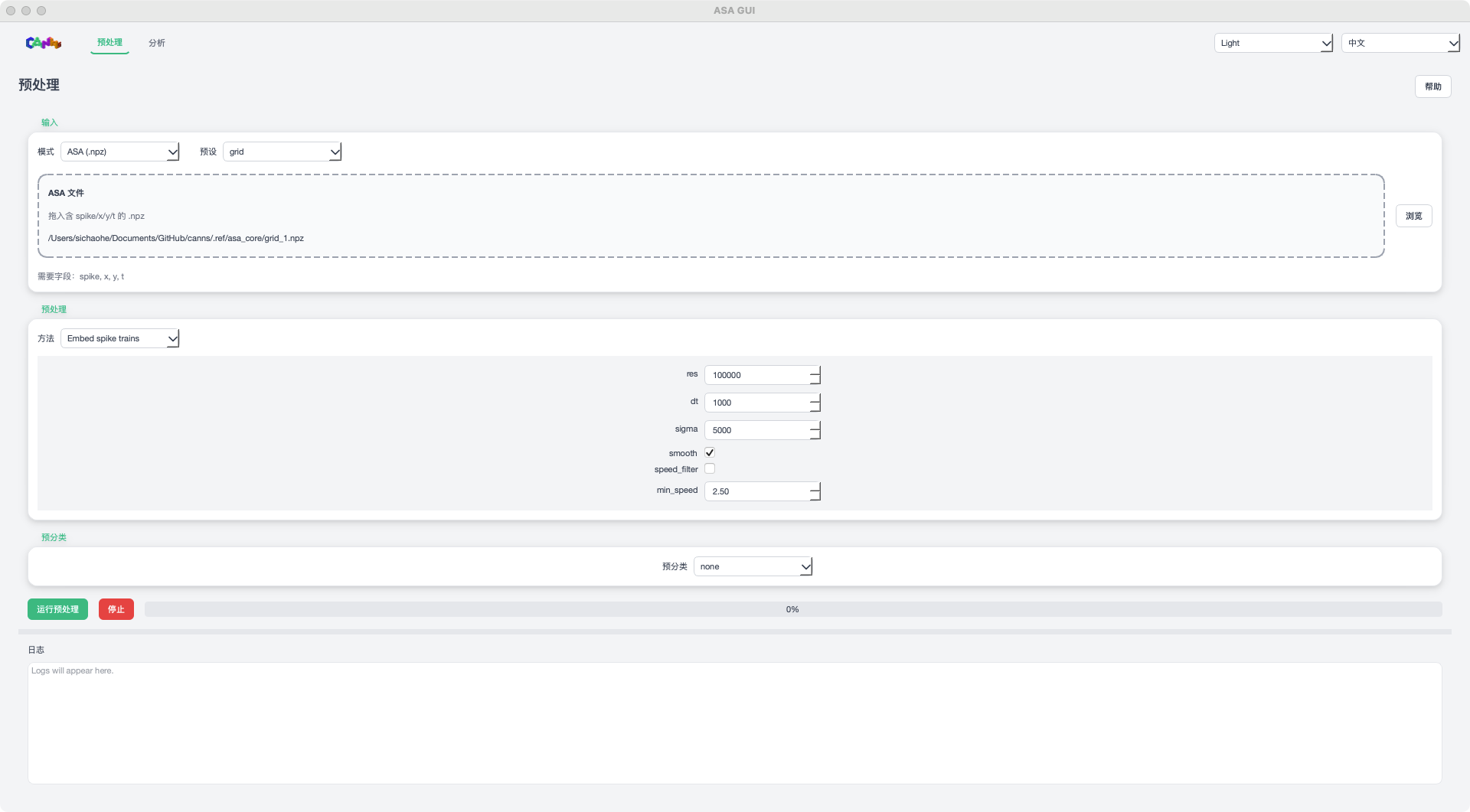
ASA GUI Preprocess page (placeholder image)¶
Analysis Page¶
Analysis Parameters |
Select analysis modules and set parameters. |
Analysis module |
Supports |
Preprocess (Standardization) |
|
Help |
Opens quick help and common tips. |
Language |
Toggle interface language between Chinese and English. |
Run Analysis / Stop / Progress |
Run analysis and show progress. |
Result Tabs |
|
Files |
Lists output files and allows Open Folder to open the results directory. |
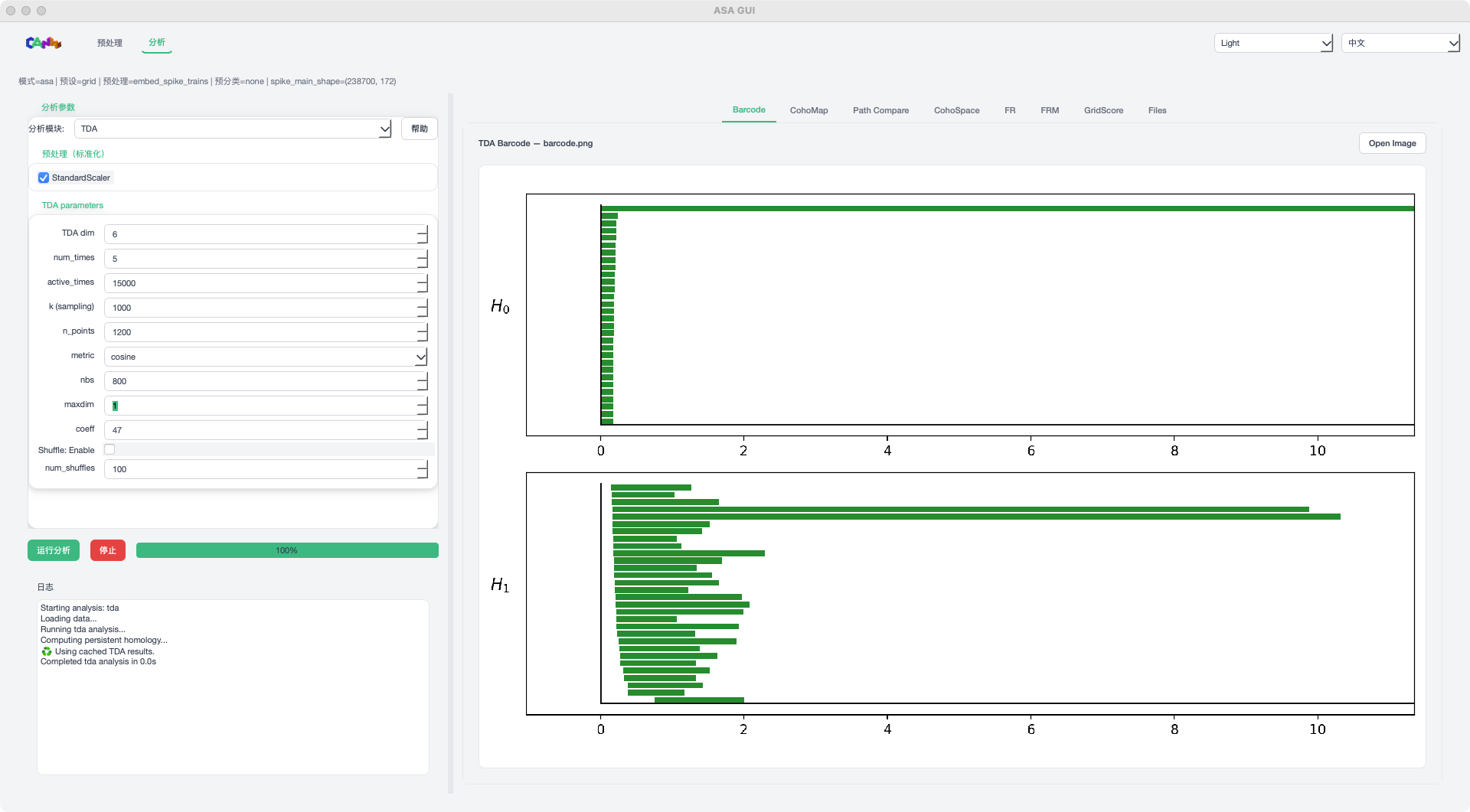
TDA mode example¶
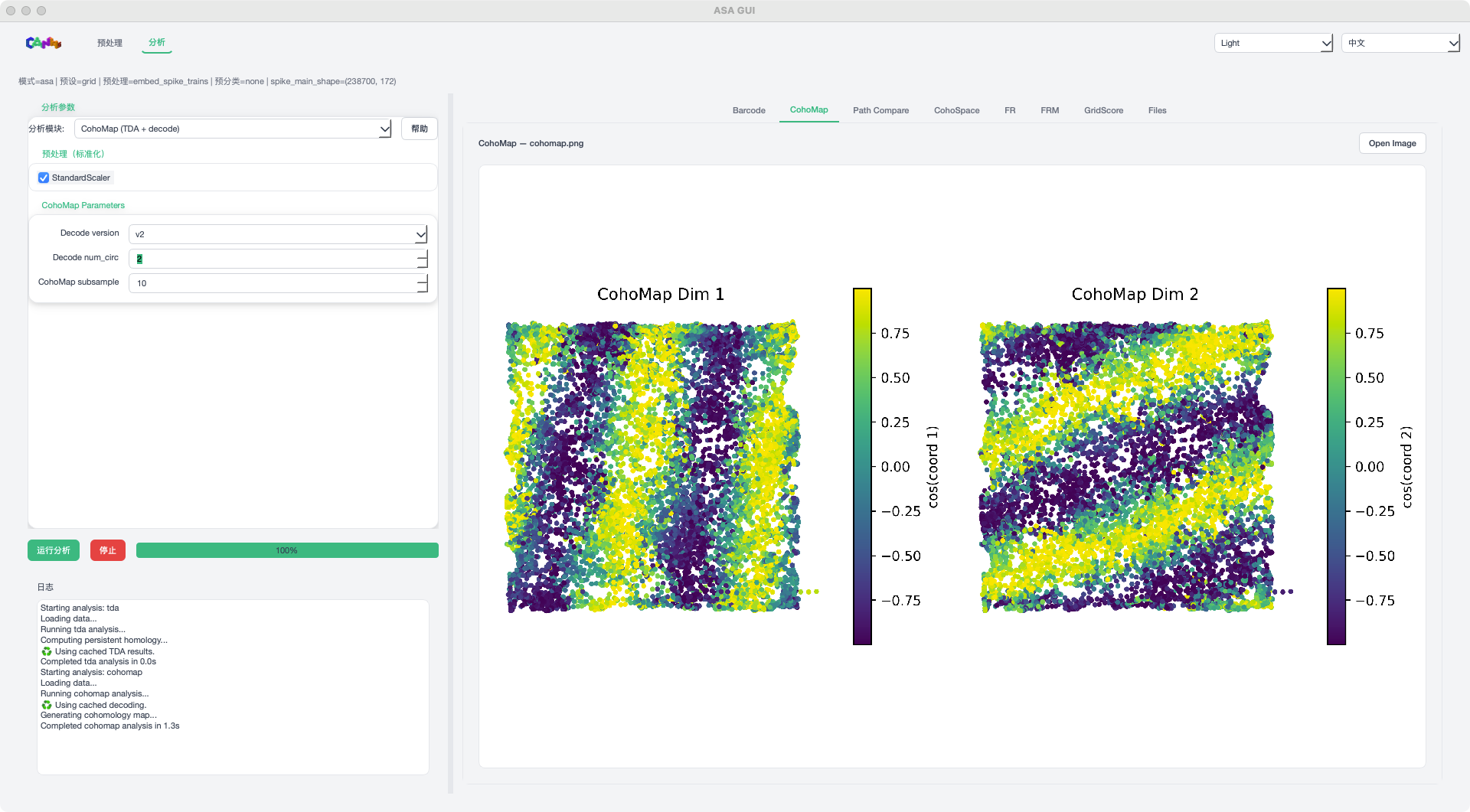
CohoMap mode example¶
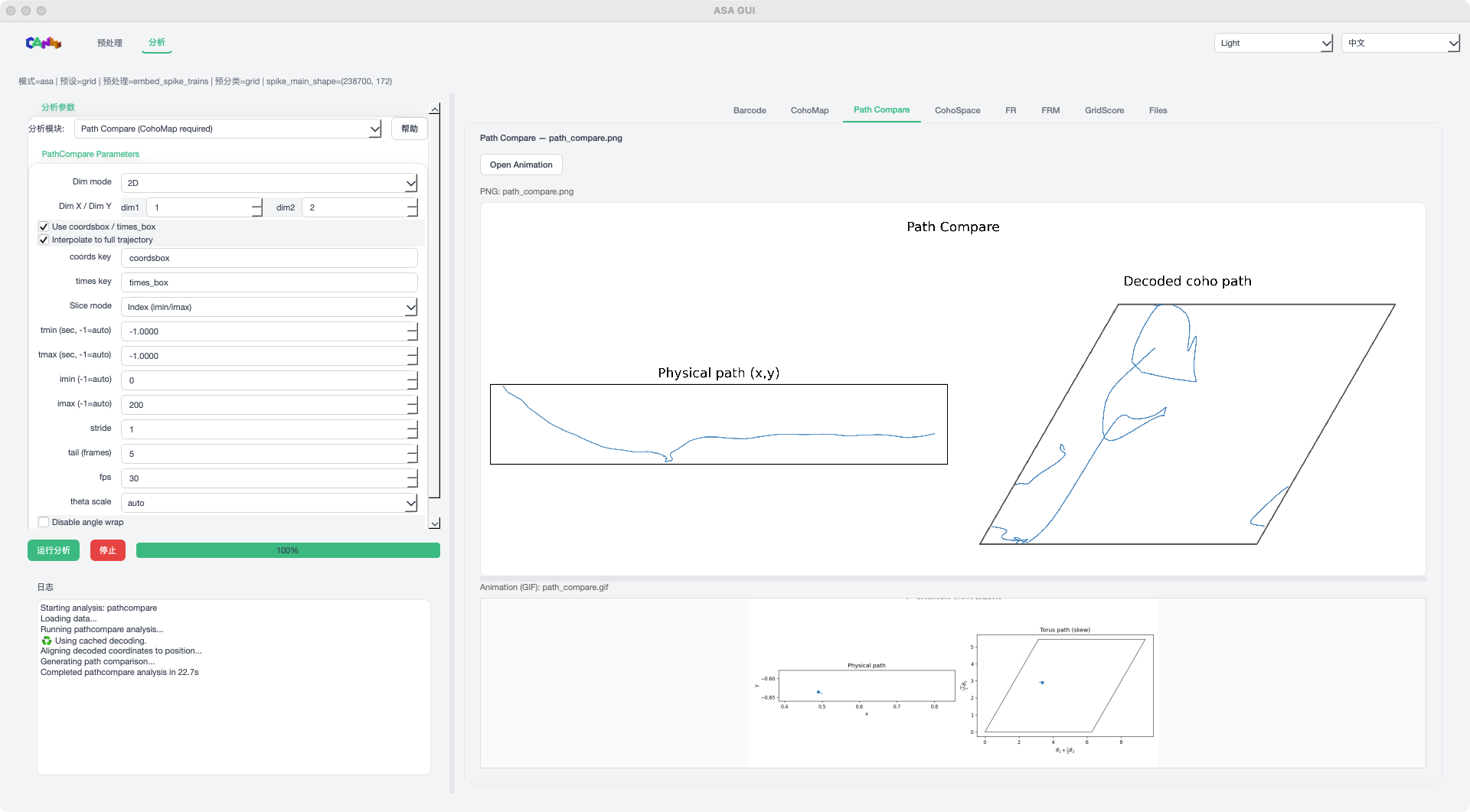
Path Compare mode example¶
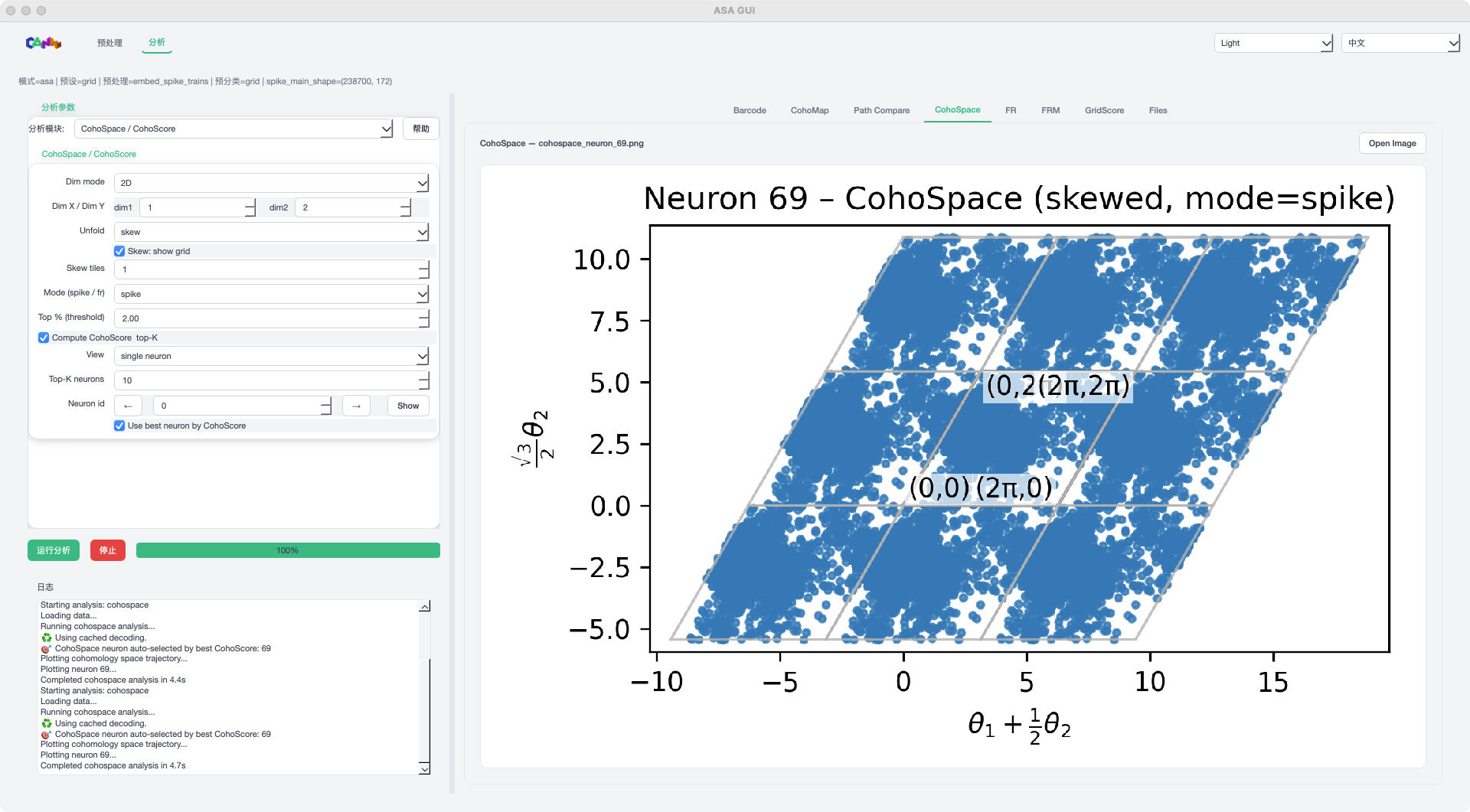
CohoSpace mode example¶
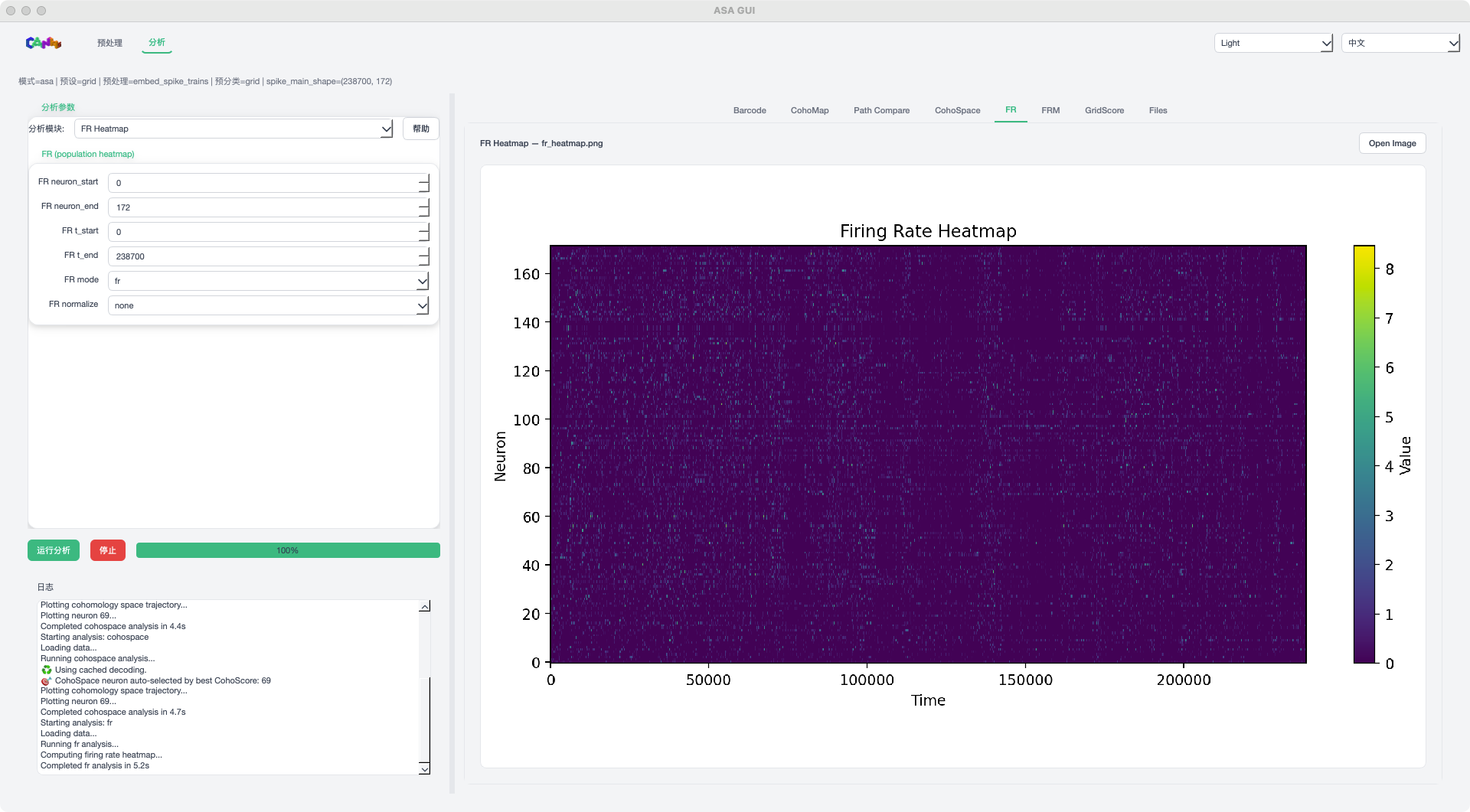
FR mode example¶
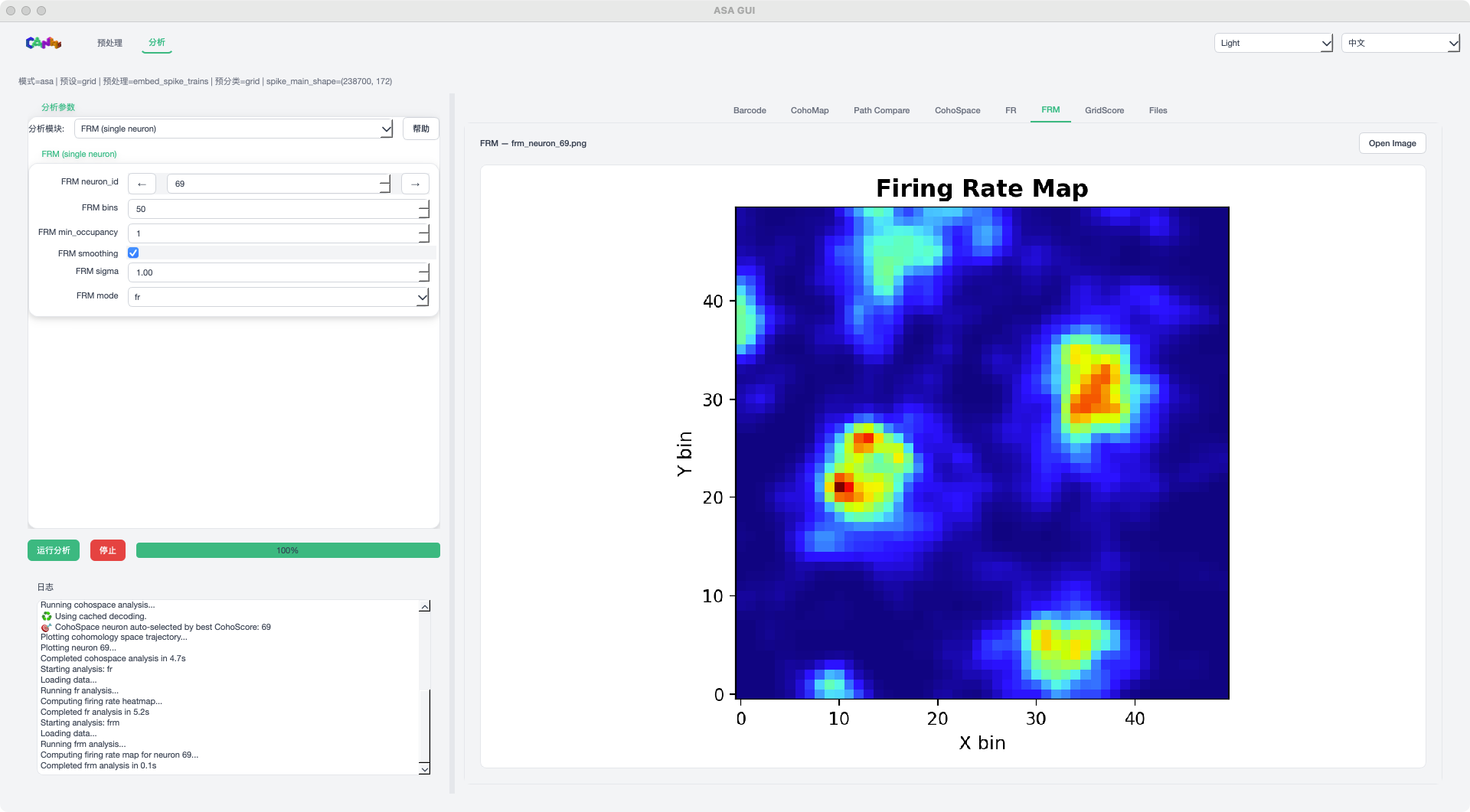
FRM mode example¶
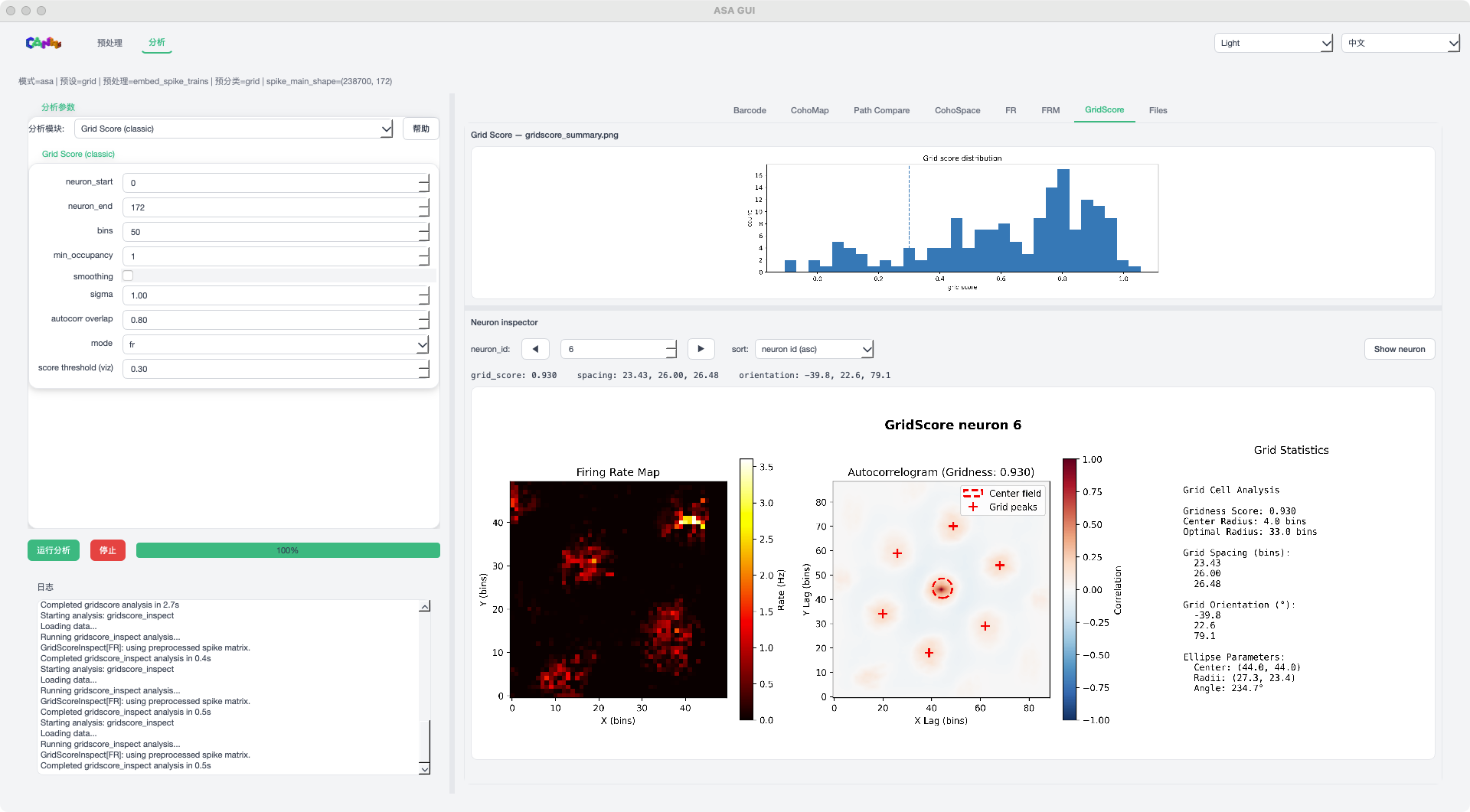
GridScore mode example¶
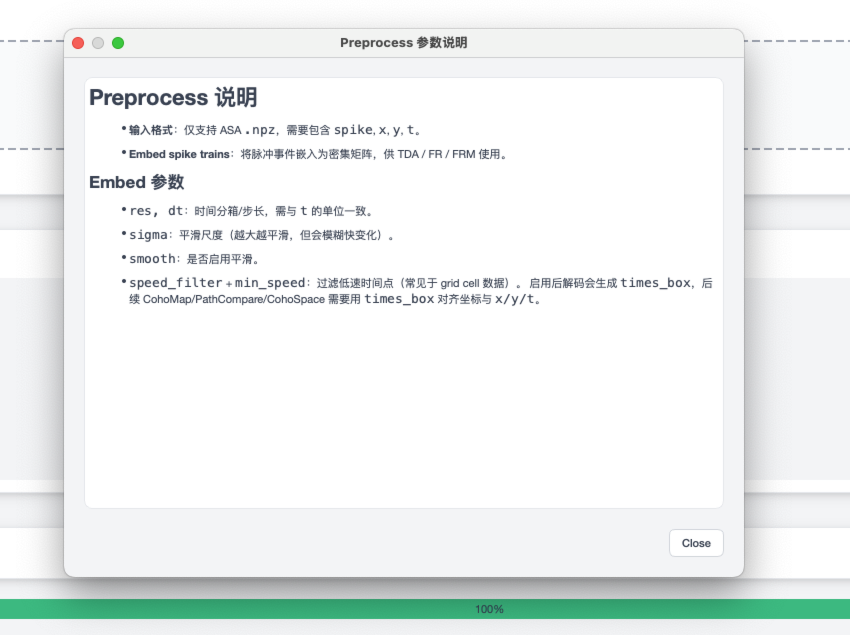
Preprocess help panel¶
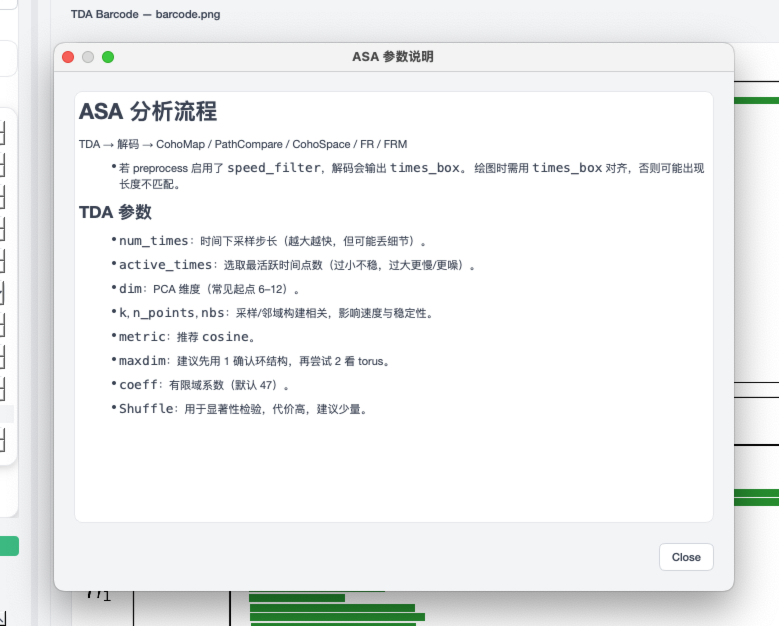
TDA help panel¶
Workflow Overview¶
Launch the ASA GUI
Select an ASA
.npzinput fileConfigure preprocessing parameters and run Preprocess
Switch to the Analysis page and select an analysis module
Run the analysis and view results in the tabs
Step 1: Select ASA File¶
In the ASA file section, drag and drop or click Browse to select a .npz file.
The interface will prompt for expected fields: spike / x / y / t.
ASA File Format¶
Must include at least spike and t:
spike: A dense matrix of shapeT x N, or a spike data structure suitable for embeddingt: Time series (synchronized withspike)Optional:
x/yfor trajectory-related analyses
import numpy as np
np.savez(
"session_asa.npz",
spike=spike,
t=t,
x=x,
y=y,
)
Step 2: Set Preprocessing¶
In the Preprocess section, choose a preprocessing method:
None: Use the original spike structure directly
Embed spike trains: Generate a dense matrix for use in TDA/FR/FRM
If embedding is required, set parameters such as res / dt / sigma / smooth /
speed_filter / min_speed, then click Run Preprocess.
Step 3: Select Analysis Module¶
Switch to the Analysis page, choose an analysis module, and configure its parameters:
TDA: Persistent homology analysis and barcode visualization
CohoMap / CohoSpace: Decode and plot spatial structures
Path Compare: Trajectory comparison (with animation output)
FR / FRM: Firing rate heatmaps and neuron firing rate maps
GridScore: Grid score computation and neuron browser
Click Run Analysis to start.
Step 4: View Results¶
After completion, view results in the right-hand tabs:
Image Tabs: Support embedded preview and Open Image to open externally
GridScore: Includes distribution plots and a neuron inspector
Files: Lists output files and allows opening the output directory
Output Directory Structure¶
The output directory defaults to the current working directory:
Results/<dataset>_<hash>/
where <dataset> is derived from the input filename and <hash> is a prefix of the input hash.
The directory contains analysis results and a cache (.asa_cache) to accelerate repeated runs.
Notes¶
The GUI’s working directory is the current directory at launch time. Change directories in the terminal before launching if needed.
For
Neuron + Trajectoryinput, please use the ASA TUI or script-based workflow instead.