Tutorial 3: Cell Classification¶
Objective: Demonstrate how to classify cells based on GridScore and firing rate statistics, and output the classification results along with visualizations.
Structure:
Task Overview
Compute GridScore
Example Classification Method
Output and Visualization
Note
This tutorial is based on the scripts in examples/cell_classification and focuses on examples and workflow.
For a complete ASA pipeline, please first read Tutorial 1: ASA Pipeline: End-to-End Decoding of Experimental Data.
0. Task Overview¶
Cell classification is typically based on the following information:
GridScore: Measures whether the firing field exhibits a grid-like structure
Firing rate and spatial selectivity: Used to filter candidate cells
Module assignment: Identifies distinct grid modules through clustering of autocorrelation features
This tutorial demonstrates two practical approaches:
Single-cell GridScore calculation and visualization
Module identification (classification) based on autocorrelation features
1. Compute GridScore¶
Example script: examples/cell_classification/grid_score.py
Key steps:
embed_spike_trainsembeds spike events into a(T, N)matrixcompute_rate_map_from_binnedgenerates the rate mapcompute_2d_autocorrelationcomputes the autocorrelationGridnessAnalyzer.compute_gridness_scoreyields the GridScore
[ ]:
from canns.analyzer import data
from canns.analyzer.data.cell_classification import (
GridnessAnalyzer,
compute_2d_autocorrelation,
compute_rate_map_from_binned,
plot_autocorrelogram,
plot_rate_map,
)
from canns.data.loaders import load_grid_data, load_left_right_npz
grid_data = load_grid_data()
cfg = data.SpikeEmbeddingConfig(smooth=False, speed_filter=True, min_speed=0.0)
spikes, xx, yy, _tt = data.embed_spike_trains(grid_data, config=cfg)
analyzer = GridnessAnalyzer()
neuron_id = 69
rate_map, *_ = compute_rate_map_from_binned(xx, yy, spikes[:, neuron_id], bins=35)
autocorr = compute_2d_autocorrelation(rate_map)
result = analyzer.compute_gridness_score(autocorr)
print(f"neuron {neuron_id}\nscore: {result.score}\nspacing: {result.spacing}\norientation: {result.orientation}")
# Visualize rate map + autocorrelogram
plot_rate_map(rate_map, show=True)
plot_autocorrelogram(autocorr, gridness_score=result.score, show=True)
Loaded grid_1: 172 neurons
Position data available: 238700 time points
neuron 69
score: 0.8761650577011183
spacing: [18.43908891 17.08800749 18.11077028]
orientation: [-40.60129465 20.55604522 83.65980825]
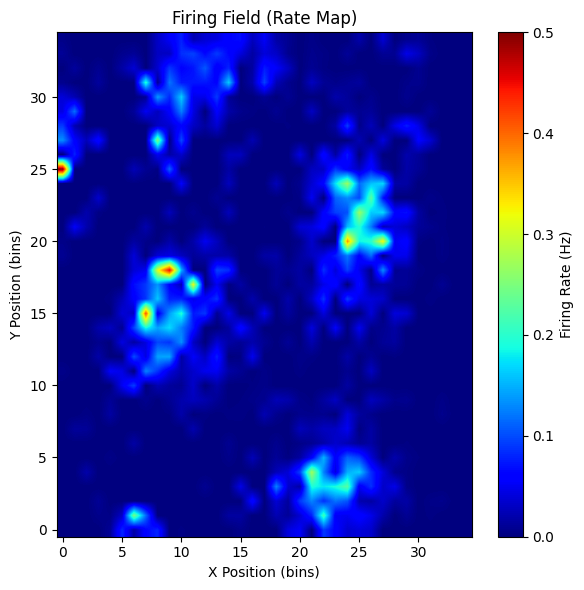
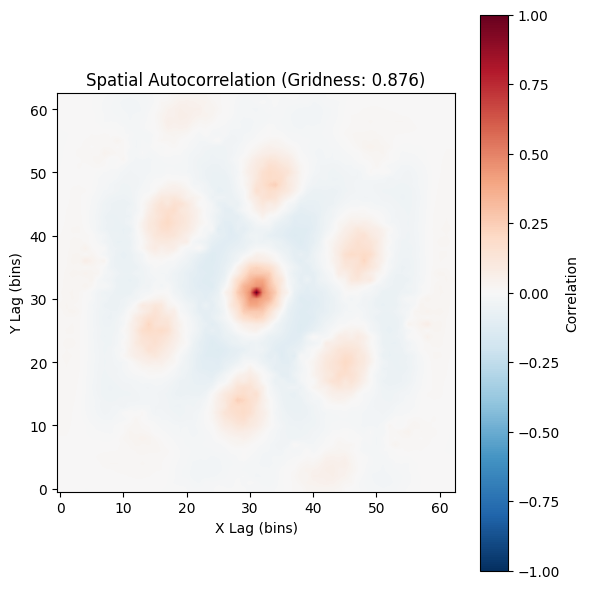
(<Figure size 600x600 with 2 Axes>,
<Axes: title={'center': 'Spatial Autocorrelation (Gridness: 0.876)'}, xlabel='X Lag (bins)', ylabel='Y Lag (bins)'>)
2. Classification Method Examples¶
Example script: examples/cell_classification/try_classification_methods.py
This example uses autocorrelograms as features and applies Leiden clustering to identify grid modules. Core function:
identify_grid_modules_and_stats: outputs module assignments and statistical information
[8]:
from canns.analyzer.data.cell_classification import (
GridnessAnalyzer,
identify_grid_modules_and_stats,
compute_rate_map_from_binned,
compute_2d_autocorrelation,
)
import numpy as np
grid_data = load_left_right_npz(
session_id="28304_1",
filename="28304_1_ASA_mec_full_cm.npz",
)
n_units = spikes.shape[1]
autocorrs = []
for nid in range(n_units):
rate_map, *_ = compute_rate_map_from_binned(xx, yy, spikes[:, nid], bins=35)
autocorr = compute_2d_autocorrelation(rate_map)
autocorrs.append(autocorr.astype(np.float32, copy=False))
autocorrs = np.stack(autocorrs, axis=0)
analyzer = GridnessAnalyzer()
out = identify_grid_modules_and_stats(
autocorrs,
gridness_analyzer=analyzer,
k=30,
resolution=1.0,
score_thr=0.3,
consistency_thr=0.5,
min_cells=10,
merge_corr_thr=0.7,
metric='manhattan',
)
print(out['n_grid_cells'], out['n_modules'])
106 1
/Users/sichaohe/Documents/GitHub/canns/src/canns/analyzer/data/cell_classification/utils/image_processing.py:284: FutureWarning: `binary_dilation` is deprecated since version 0.26 and will be removed in version 0.28. Use `skimage.morphology.dilation` instead. Note the lack of mirroring for non-symmetric footprints (see docstring notes).
dilated = morphology.binary_dilation(image, footprint=footprint)
3. Output and Visualization¶
grid_score.pywill plot the rate map, autocorrelation map, and GridScore histogram.try_classification_methods.pywill savegrid_modules_demo_output.npz.
Recommendations:
First validate parameters using a small batch of neurons before scaling up to the entire population.
Adjust
bins/score_thr/consistency_thrappropriately to match data quality.If integration with the ASA pipeline is required, combine
spikesandx/y/twith decoding results for joint analysis.